A guided pipeline for Aligning Patients to Cas
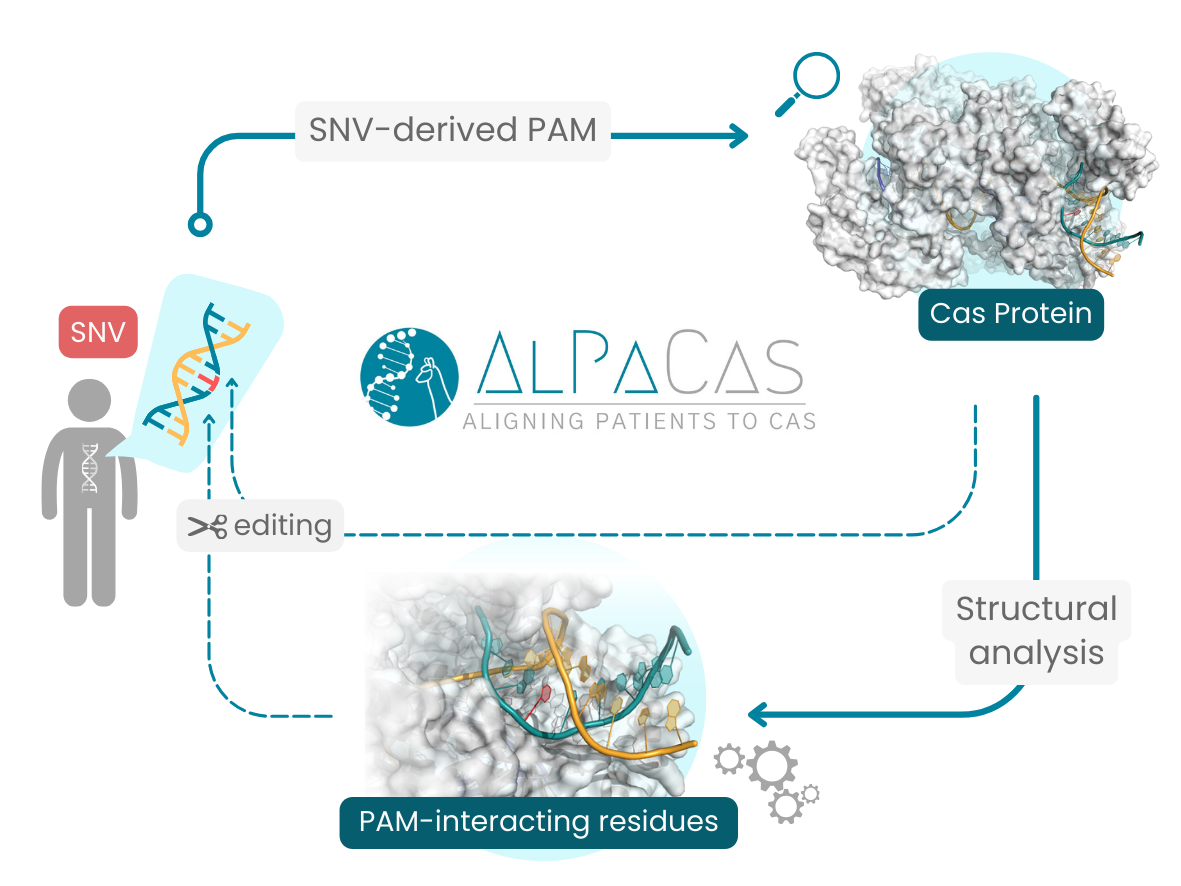
Engineering the Cas-PAM interaction has emerged as an effective protocol for achieving allele-specific targeting, whereas the permissive nature of DNA-RNA interactions alone would not have been sufficient. Such SNV-derived PAM approaches are intended to exploit the formation of PAM sequence solely on the mutated allele, discarding the wild-type allele from cleavage.
In this context, we have developed AlPaCas, to streamline the identification and analysis of SNV-derived PAMs.
With AlPaCas you can:
- Analyze ClinVar-annotated SNVs
- Run structural analyses in the context of selected SNV-derived PAMs
- Customize your search by adding your SNVs, PAM patterns, or structures.
... and more. See the HELP page for further details.



NAR Web Server Issue 2024 DOI Serena Rosignoli, Elisa Lustrino, Alessio Conci, Alessandra Fabrizi, Serena Rinaldo, Maria Carmela Latella, Elena Enzo, Gianni Prosseda, Laura De Rosa, Michele De Luca, Alessandro Paiardini, AlPaCas: allele-specific CRISPR gene editing through a protospacer adjacent motif (PAM) approach , Nucleic Acids Research, 2024
Last databases update: 2024-07-23 12:02:30
This website is free, open to all users, there is no login requirement and doesn't use cookies.
This work is licensed under CC BY-NC 4.0



